Genetic 564 Undergraduate Capstone course at University of Wisconsin Madison
What is Homology?
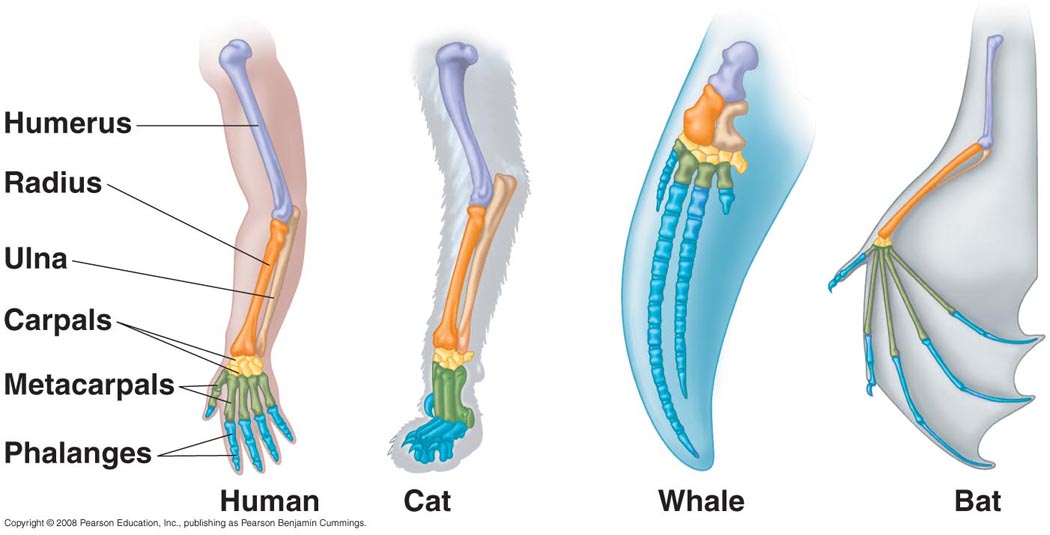
Homology refers to an object having a similar or same relationship, structure, or location. [1] In the field of genomics when referring to homology it is associated how similar/dissimilar DNA sequences from two or more organisms, or in proteomics protein sequences. [2] Homology is an evolutionary tool that provides historic context, for example investigating biological structures. Fig.1
Fig. 1 Homology of human arm compared to other mammals. All four organisms share the same bone number and placement, however the structure and functions differ.
Homology of PAX8
Human (H. sapien) and many other mammals can suffer from congenital hypothyroidism that is linked to PAX8 mutations. The gene PAX8 itself can be located in vertebrate genomes, mainly to mammals, however homologs of PAX8 exist in amphibians and fish. The fruit fly (Drosophila melanogaster) being an invertebrate organism with a divergent endocrine system from vertebrates contains ancestral portions of the PAX8 known as "shaven" (dPAX2) which transcribes different physiological functions.


Species: P.troglodytes
Gene: PAX8 FASTA: XP_003309249.2 Percent Identity: 94% Amino Acid Length: 389 


|





Species: D. melanogaster
Gene: shaven [dPAX2] FASTA: FBgn0005561 Percent Identity: 22% Amino Acid Length: 844 |
Importance of PAX8 Homology
Having protein and gene homology is crucial when considering experimental design in which the researcher when selecting for a model organism. Conservation of function and sequence is an essential component to consider in the context of evolution over time. When comparing PAX8 homology between D. melanogaster (Fruit Flies) to other mammals not only is the sequence length significantly different, and serve different functions. Resulting in usage of Fruit Flies for PAX8 or in endocrine research not recommended. However, comparing mammals and human homology is supported by high percent identity, and similar amino acid (aa) length can be used to support decision in regards to model organism selection.
References:
[1]Pinna, M. C. (1991). Concepts And Tests Of Homology In The Cladistic Paradigm. Cladistics, 7(4), 367-394. doi:10.1111/j.1096-0031.1991.tb00045.x
[2]Wagner, G. P. (2007). The developmental genetics of homology. Nature Reviews Genetics, 8(6), 473-479. doi:10.1038/nrg2099
Images:
Cover photo: https://www.wddty.com/img/wddty/magazine/large/4643.jpg
[1]Pinna, M. C. (1991). Concepts And Tests Of Homology In The Cladistic Paradigm. Cladistics, 7(4), 367-394. doi:10.1111/j.1096-0031.1991.tb00045.x
[2]Wagner, G. P. (2007). The developmental genetics of homology. Nature Reviews Genetics, 8(6), 473-479. doi:10.1038/nrg2099
Images:
Cover photo: https://www.wddty.com/img/wddty/magazine/large/4643.jpg